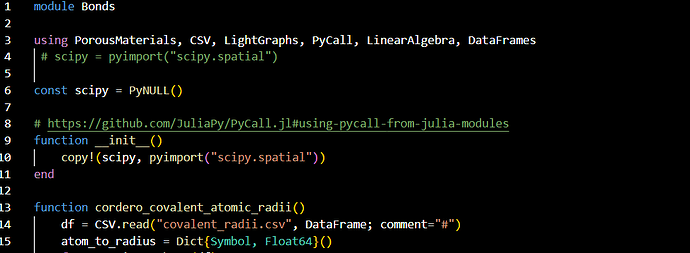
在Bonds.jl里,加个using DataFrame?
![]()
可以,最好把其它也放进去,就不用每次打这么多字了
julia> push!(LOAD_PATH,“D:/julia”)
4-element Vector{String}:
“@”
“@v#.#”
“@stdlib”
“D:/julia”
julia> using Bonds
[ Info: Precompiling Bonds [top-level]
┌ Warning: crystals path directory not found
│ path = “C:\Users\zzh666\data\crystals”
└ @ Xtals C:\Users\zzh666.julia\packages\Xtals\vIP0H\src\misc.jl:186
┌ Warning: data path directory not found
│ path = “C:\Users\zzh666\data”
└ @ Xtals C:\Users\zzh666.julia\packages\Xtals\vIP0H\src\misc.jl:186
┌ Warning: forcefields path directory not found
│ path = “C:\Users\zzh666\data\forcefields”
└ @ Xtals C:\Users\zzh666.julia\packages\Xtals\vIP0H\src\misc.jl:186
┌ Warning: grids path directory not found
│ path = “C:\Users\zzh666\data\grids”
└ @ Xtals C:\Users\zzh666.julia\packages\Xtals\vIP0H\src\misc.jl:186
┌ Warning: molecules path directory not found
│ path = “C:\Users\zzh666\data\molecules”
└ @ Xtals C:\Users\zzh666.julia\packages\Xtals\vIP0H\src\misc.jl:186
┌ Warning: crystals path directory not found
│ path = “C:\Users\zzh666\data\crystals”
└ @ Xtals C:\Users\zzh666.julia\packages\Xtals\vIP0H\src\misc.jl:186
┌ Warning: data path directory not found
│ path = “C:\Users\zzh666\data”
└ @ Xtals C:\Users\zzh666.julia\packages\Xtals\vIP0H\src\misc.jl:186
┌ Warning: simulations path directory not found
│ path = “C:\Users\zzh666\data\simulations”
└ @ Xtals C:\Users\zzh666.julia\packages\Xtals\vIP0H\src\misc.jl:186
ERROR: LoadError: ArgumentError: Package DataFrame not found in current path:
- Run
import Pkg; Pkg.add("DataFrame")to install the DataFrame package.
Stacktrace:
[1] require(into::Module, mod::Symbol)
@ Base .\loading.jl:893
[2] include
@ .\Base.jl:384 [inlined]
[3] include_package_for_output(pkg::Base.PkgId, input::String, depot_path::Vector{String}, dl_load_path::Vector{String}, load_path::Vector{String}, concrete_deps::Vector{Pair{Base.PkgId, UInt64}}, source::Nothing)
@ Base .\loading.jl:1235
[4] top-level scope
@ none:1
[5] eval
@ .\boot.jl:360 [inlined]
[6] eval(x::Expr)
@ Base.MainInclude .\client.jl:446
[7] top-level scope
@ none:1
in expression starting at D:\julia\Bonds.jl:1
ERROR: Failed to precompile Bonds [top-level] to C:\Users\zzh666.julia\compiled\v1.6\jl_B8BA.tmp.
Stacktrace:
[1] error(s::String)
@ Base .\error.jl:33
[2] compilecache(pkg::Base.PkgId, path::String, internal_stderr::Base.TTY, internal_stdout::Base.TTY, ignore_loaded_modules::Bool)
@ Base .\loading.jl:1385
[3] compilecache(pkg::Base.PkgId, path::String)
@ Base .\loading.jl:1329
[4] _require(pkg::Base.PkgId)
@ Base .\loading.jl:1043
[5] require(uuidkey::Base.PkgId)
@ Base .\loading.jl:936
[6] require(into::Module, mod::Symbol)
@ Base .\loading.jl:923
又说Package DataFrame not found in current path:
有没有一种可能,这个包叫DataFrames
你说的对 ![]()
但在cmd里新问题:
ERROR: LoadError: ArgumentError: “covalent_radii.csv” is not a valid file
Stacktrace:
[1] Header
@ C:\Users\zzh666.julia\packages\CSV\Zl2ww\src\header.jl:90 [inlined]
[2] CSV.File(source::String; header::Int64, normalizenames::Bool, datarow::Int64, skipto::Nothing, footerskip::Int64, transpose::Bool, comment::String, use_mmap::Nothing, ignoreemptylines::Bool, select::Nothing, drop::Nothing, missingstrings::Vector{String}, missingstring::String, delim::Nothing, ignorerepeated::Bool, quotechar::Char, openquotechar::Nothing, closequotechar::Nothing, escapechar::Char, dateformat::Nothing, dateformats::Nothing, decimal::UInt8, truestrings::Vector{String}, falsestrings::Vector{String}, type::Nothing, types::Nothing, typemap::Dict{Type, Type}, pool::Float64, lazystrings::Bool, strict::Bool, silencewarnings::Bool, debug::Bool, parsingdebug::Bool, kw::Base.Iterators.Pairs{Union{}, Union{}, Tuple{}, NamedTuple{(), Tuple{}}})
@ CSV C:\Users\zzh666.julia\packages\CSV\Zl2ww\src\file.jl:223
[3] read(source::String, sink::Type; copycols::Bool, kwargs::Base.Iterators.Pairs{Symbol, String, Tuple{Symbol}, NamedTuple{(:comment,), Tuple{String}}})
@ CSV C:\Users\zzh666.julia\packages\CSV\Zl2ww\src\CSV.jl:45
[4] cordero_covalent_atomic_radii()
@ Bonds D:\julia\Bonds.jl:14
[5] top-level scope
@ D:\julia\Bonds.jl:34
[6] include
@ .\Base.jl:384 [inlined]
[7] include_package_for_output(pkg::Base.PkgId, input::String, depot_path::Vector{String}, dl_load_path::Vector{String}, load_path::Vector{String}, concrete_deps::Vector{Pair{Base.PkgId, UInt64}}, source::Nothing)
@ Base .\loading.jl:1235
[8] top-level scope
@ none:1
[9] eval
@ .\boot.jl:360 [inlined]
[10] eval(x::Expr)
@ Base.MainInclude .\client.jl:446
[11] top-level scope
@ none:1
in expression starting at D:\julia\Bonds.jl:1
ERROR: Failed to precompile Bonds [top-level] to C:\Users\zzh666.julia\compiled\v1.6\jl_D354.tmp.
Stacktrace:
[1] error(s::String)
@ Base .\error.jl:33
[2] compilecache(pkg::Base.PkgId, path::String, internal_stderr::Base.TTY, internal_stdout::Base.TTY, ignore_loaded_modules::Bool)
@ Base .\loading.jl:1385
[3] compilecache(pkg::Base.PkgId, path::String)
@ Base .\loading.jl:1329
[4] _require(pkg::Base.PkgId)
@ Base .\loading.jl:1043
[5] require(uuidkey::Base.PkgId)
@ Base .\loading.jl:936
[6] require(into::Module, mod::Symbol)
@ Base .\loading.jl:923
用cd切换当前目录:
cd("D:/julia")
什么cd,在哪,我没找到cd ![]()
这样吗,将“covalent_radii.csv”路径改为当前路径
ERROR: InitError: PyError (PyImport_ImportModule
The Python package scipy.spatial could not be imported by pyimport. Usually this means
that you did not install scipy.spatial in the Python version being used by PyCall.
PyCall is currently configured to use the Julia-specific Python distribution
installed by the Conda.jl package. To install the scipy.spatial module, you can
use pyimport_conda("scipy.spatial", PKG), where PKG is the Anaconda
package that contains the module scipy.spatial, or alternatively you can use the
Conda package directly (via using Conda followed by Conda.add etcetera).
Alternatively, if you want to use a different Python distribution on your
system, such as a system-wide Python (as opposed to the Julia-specific Python),
you can re-configure PyCall with that Python. As explained in the PyCall
documentation, set ENV[“PYTHON”] to the path/name of the python executable
you want to use, run Pkg.build(“PyCall”), and re-launch Julia.
) <class ‘ModuleNotFoundError’>
ModuleNotFoundError(“No module named ‘scipy’”)
Stacktrace:
[1] pyimport(name::String)
@ PyCall C:\Users\zzh666.julia\packages\PyCall\7a7w0\src\PyCall.jl:550
[2] init()
@ Bonds D:\julia\Bonds.jl:10
[3] _include_from_serialized(path::String, depmods::Vector{Any})
@ Base .\loading.jl:696
[4] _require_from_serialized(path::String)
@ Base .\loading.jl:749
[5] _require(pkg::Base.PkgId)
@ Base .\loading.jl:1053
[6] require(uuidkey::Base.PkgId)
@ Base .\loading.jl:936
[7] require(into::Module, mod::Symbol)
@ Base .\loading.jl:923
during initialization of module Bonds
然后这样的
可以,但是得记得每个都要改
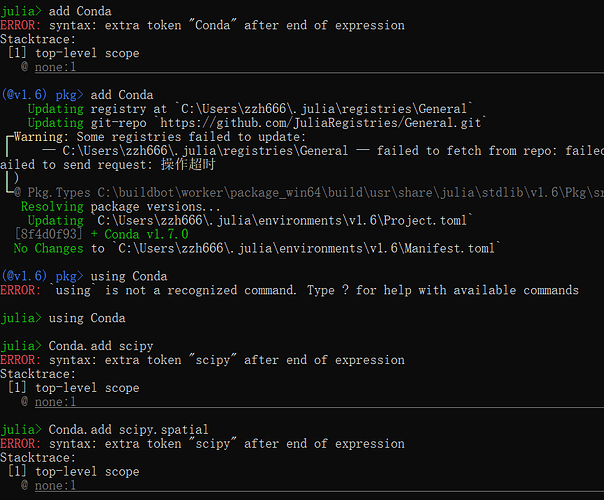
julia> Conda.add(“scipy.spatial”)
[ Info: Running conda install -y scipy.spatial in root environment
Collecting package metadata (current_repodata.json): done
Solving environment: failed with initial frozen solve. Retrying with flexible solve.
Collecting package metadata (repodata.json): done
Solving environment: failed with initial frozen solve. Retrying with flexible solve.
PackagesNotFoundError: The following packages are not available from current channels:
- scipy.spatial
Current channels:
- conda-forge/win-64
- conda-forge/noarch
- main/win-64
- main/noarch
- r/win-64
- r/noarch
- msys2/win-64
- msys2/noarch
To search for alternate channels that may provide the conda package you’re
looking for, navigate to
https://anaconda.org
and use the search bar at the top of the page.
ERROR: failed process: Process(setenv('C:\Users\zzh666\.julia\conda\3\Scripts\conda.exe' install -y scipy.spatial,[“PATH=C:\Users\zzh666\.julia\conda\3\Library\bin;C:\Program Files (x86)\Intel\Intel(R) Management Engine Components\iCLS\;C:\Program Files\Intel\Intel(R) Management Engine Components\iCLS\;C:\Windows\system32;C:\Windows;C:\Windows\System32\Wbem;C:\Windows\System32\WindowsPowerShell\v1.0\;C:\Windows\System32\OpenSSH\;C:\Program Files (x86)\Intel\Intel(R) Management Engine Components\DAL;C:\Program Files\Intel\Intel(R) Management Engine Components\DAL;C:\Program Files (x86)\Intel\Intel(R) Management Engine Components\IPT;C:\Program Files\Intel\Intel(R) Management Engine Components\IPT;C:\Program Files (x86)\NVIDIA Corporation\PhysX\Common;C:\WINDOWS\system32;C:\WINDOWS;C:\WINDOWS\System32\Wbem;C:\WINDOWS\System32\WindowsPowerShell\v1.0\;C:\WINDOWS\System32\OpenSSH\;D:\nanconda3;D:\nanconda3\Scripts;D:\julia\Julia-1.3.1\bin\julia.exe;C:\Users\zzh666\AppData\Local\Microsoft\WindowsApps;;D:\pycharm\PyCharm Community Edition 2021.3.3\bin;;D:\Julia\Julia-1.7.3\bin;F:\VScode\Microsoft VS Code\bin;D:\julia\Julia-1.6.6\bin”, “USERDOMAIN_ROAMINGPROFILE=LAPTOP-GGS2MDC7”, “HOMEPATH=\Users\zzh666”, “PATHEXT=.COM;.EXE;.BAT;.CMD;.VBS;.VBE;.JS;.JSE;.WSF;.WSH;.MSC”, “SESSIONNAME=Console”, “SYSTEMROOT=C:\WINDOWS”, “APPDATA=C:\Users\zzh666\AppData\Roaming”, “PSMODULEPATH=C:\Program Files\WindowsPowerShell\Modules;C:\WINDOWS\system32\WindowsPowerShell\v1.0\Modules”, “COMMONPROGRAMW6432=C:\Program Files\Common Files”, “PROGRAMDATA=C:\ProgramData” … “PROGRAMFILES(X86)=C:\Program Files (x86)”, “JUNORC_PATH=F:\Juliapro\.atom”, “PROGRAMFILES=C:\Program Files”, “LOGONSERVER=\\LAPTOP-GGS2MDC7”, “DRIVERDATA=C:\Windows\System32\Drivers\DriverData”, “CONDA_PREFIX=C:\Users\zzh666\.julia\conda\3”, “SYSTEMDRIVE=C:”, “PYCHARM COMMUNITY EDITION=D:\pycharm\PyCharm Community Edition 2021.3.3\bin;”, “PROCESSOR_ARCHITECTURE=AMD64”, “OPENBLAS_MAIN_FREE=1”]), ProcessExited(1)) [1]
Stacktrace:
[1] pipeline_error
@ .\process.jl:538 [inlined]
[2] run(::Cmd; wait::Bool)
@ Base .\process.jl:453
[3] run
@ .\process.jl:451 [inlined]
[4] runconda(args::Cmd, env::String)
@ Conda C:\Users\zzh666.julia\packages\Conda\x2UxR\src\Conda.jl:128
[5] add(pkg::String, env::String; channel::String)
@ Conda C:\Users\zzh666.julia\packages\Conda\x2UxR\src\Conda.jl:222
[6] add (repeats 2 times)
@ C:\Users\zzh666.julia\packages\Conda\x2UxR\src\Conda.jl:221 [inlined]
[7] top-level scope
@ REPL[9]:1
不知到为什么会这样
抱歉,多有打扰,感觉应该快解决了,谢谢一直帮我解决问题的好心大佬了
是的,我在python都是 pip install 某模块。意思是pip install scipy.spatial吗
不要泄露个人信息;微信不能用markdown
discourse有私信功能
我的天那,你们还是去QQ上聊吧 ![]()